We present CaMol, a novel architecture for predicting molecular property in few-shot scenarios and developed by NS Lab, CUK based on pure PyTorch backend.
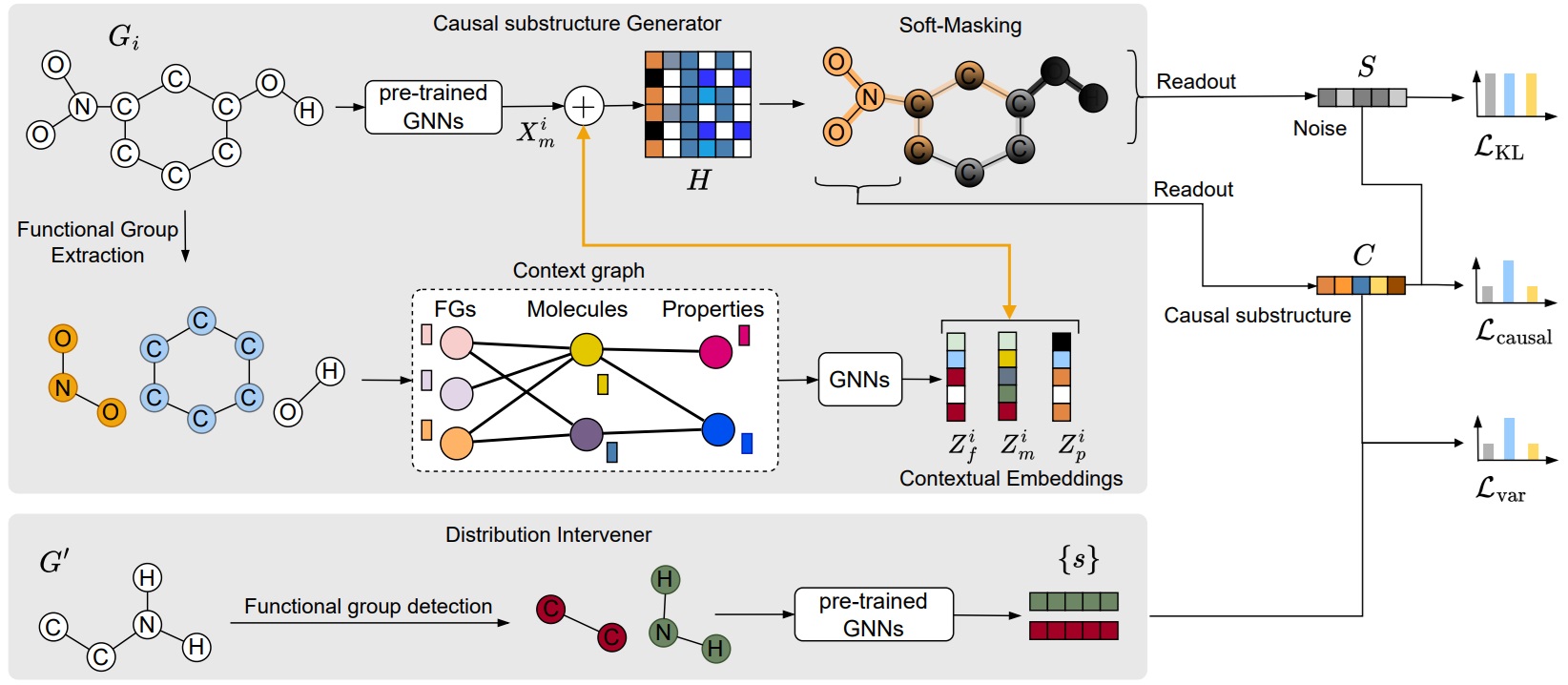
The overall architecture of CaMol.
We aim to build a context-aware graph causality inference framework to address the few-shot molecular property prediction tasks. Molecular property prediction is becoming one of the major applications of graph learning in Web-based services, e.g., online protein structure prediction and drug discovery. A key challenge arises in few-shot scenarios, where only a few labeled molecules are available for predicting unseen properties. Recently, several studies have used in-context learning to capture relationships among molecules and properties, but they face two limitations in: (1) exploiting prior knowledge of functional groups that are causally linked to properties and (2) identifying key substructures directly correlated with properties. We propose CaMol, a context-aware graph causality inference framework, to address these challenges by using a causal inference perspective, assuming that each molecule consists of a latent causal structure that determines a specific property. First, we introduce a context graph that encodes chemical knowledge by linking functional groups, molecules, and properties to guide the discovery of causal substructures. Second, we propose a learnable atom soft-masking strategy to disentangle causal substructures from confounding ones. Third, we introduce a distribution intervener that applies backdoor adjustment by combining causal substructures with chemically grounded confounders, disentangling causal effects from real-world chemical variations. Experiments on diverse molecular datasets showed that CaMol achieved superior accuracy and sample efficiency in few-shot tasks, showing its generalizability to unseen properties. Also, the discovered causal substructures were strongly aligned with chemical knowledge about functional groups, supporting the model interpretability.
A short description of CaMol:
- The idea is based on a causal inference perspective, assuming that each molecule consists of a latent causal structure that determines a specific property.
- We introduce a context graph that encodes chemical knowledge by linking functional groups, molecules, and properties to guide the discovery of causal substructures. Second, we propose a learnable atom soft-masking strategy to disentangle causal substructures from confounding ones.
- We introduce a distribution intervener that applies backdoor adjustment by combining causal substructures with chemically grounded confounders, disentangling causal effects from real-world chemical variations.
The CaMol is available at:
Cite “CaMol” as:
Please cite our paper if you find CaMol useful in your work:
@misc{hoang2026contextawaregraphcausalityinference,
title={Context-aware Graph Causality Inference for Few-Shot Molecular Property Prediction},
author={Van Thuy Hoang and O-Joun Lee},
year={2026},
eprint={2601.11135},
archivePrefix={arXiv},
primaryClass={cs.LG},
url={https://arxiv.org/abs/2601.11135},
}
Please take a look at our unified graph transformer model, UGT, which can preserve local and globl graph structure, and community-aware graph transformer model, CGT, which can mitigate degree bias problem of message passing mechanism, and S-CGIB, which builds a pre-trained Graph Neural Network (GNN) model on molecules without human annotations or prior knowledge, together.



